Comprehensive N-glycan Characterization of USP mAb 002 Reference Standard

July 12, 2023
Monoclonal antibodies (mAbs) are synthetic proteins that mimic natural human antibodies in the immune system. They have become a valuable biologic to treat a diverse range of illnesses including cancer, cardiovascular diseases, autoimmune disorders, etc. The safety and efficacy of the mAb-based drugs rely heavily on their glycosylation pattern because the N-glycans play critical roles in important biological processes such as protein binding, stability, folding, immunogenicity, and turnover. For instance,
The changes in the terminal galactosylation and sialylation affect the half-life and the efficacy (anti-inflammatory effect) of therapeutic glycoproteins.
Afucosylated Fc N-glycans exhibit stronger ADCC and higher efficacy at lower concentrations compared to the fucosylated N-glycans.
Therefore, glycosylation is considered as Critical Quality Attributes (CQA) of therapeutic glycoproteins.
The United States Pharmacopeia (USP) has released three mAb reference standards, (i.e., USP mAb 001, USP mAb 002, and USP mAb 003) that can work as controls to help establish an efficient glycan characterization protocol. However, the biotherapeutic industry is in a dire need of an efficient and robust informatic support to unfold the heterogenous glycan structures from complex LC-MS and MS/MS spectra. In a recent experiment, GlycanExplorer™ software was used for high throughput profiling and mapping of USP mAb 002 N-glycans.
The USP mAb 002 standard was digested by PNGaseF to produce N-glycosylamines, which were rapidly labeled with Glycoworks® RapiFluor-MS (RFMS) reagent. The RFMS-labeled N-glycan samples were purified, enriched, and separated using a Waters ACQUITY UPLC Glycan BEH Amide column (2.1 × 150 mm, 1.7 µm, 130 Å) coupled with a Thermo Scientific™ VanquishTM UHPLC and an Orbitrap Elite™ mass spectrometer. For detailed structural elucidation of glycans, LC-MS/MS experiment was conducted using top 3 data-dependent acquisition in positive ion mode. GlycanExplorer™ version 1.10 was used to create a custom glycan database, perform LC-MS peak processing, generate a candidate peak list, and subject it to MS/MS database search for confident identification of glycans.
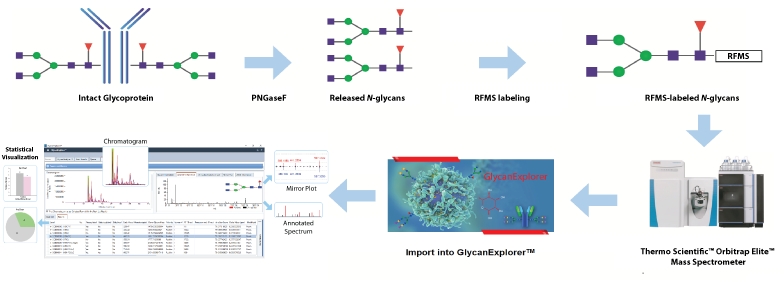
Fig: Schematic N-glycan analysis workflow
Peak Processing and MS/MS Database Search Parameters
MS1 Peak threshold intensity (with respect to the base peak): >= 0.01%
Minimum peak height (absolute XIC intensity): 2000
Chromatographic peak width: 0.1 - 1.5 mins
Range of isotopologues to validate an LC compound: [M+1] to [M+6]
Compound charge state range: 1 - 3
Fragments for MS/MS database search: Glycosidic and cross ring
Ion species: [M+H]+, [M+2H]2+, and [M+3H]3+
Custom glycan groups: Fucosylated, Galactosylated, and Sialylated
GlycanExplorer software detected 10,497 LC compounds and annotated 22 unique glycan structures. F(6)A2 was found to be the most abundant glycan with a relative intensity of 54.09% and M7D3 is the lowest abundant one (0.034%). Of all the 22 identified glycans, 94.52% are complex N-glycans and 4.58% are oligomannose structures. The N-glycan profile of the USP mAb 002 reference standard reveals that 94.52% (15) of the N-glycans are fucosylated, 27.23% (9) are galactosylated, and 1.72% (3) are sialylated.
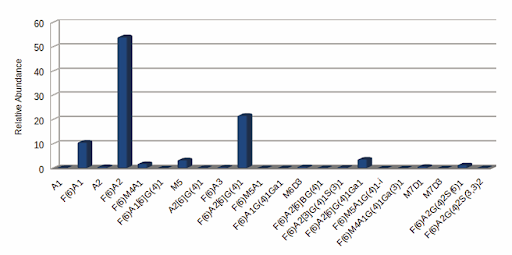
Fig: USP mAb 002 N-glycan expression plot
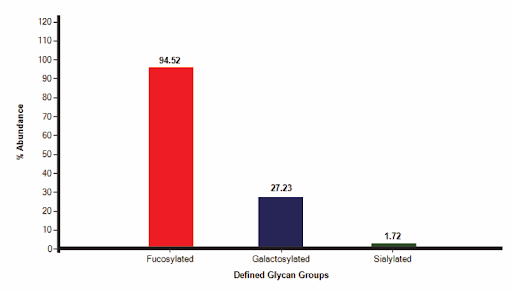
Fig: N-glycosylation profile of USP mAb 002 reference standard
| Comment | Share |
|


