Home >> Products >> MALDIVision
A comprehensive data processing & visualization tool for MALDI Imaging Mass Spectrometric data
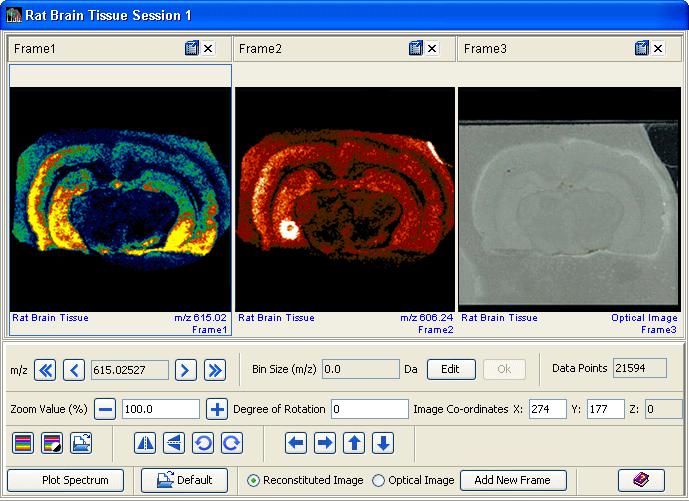
MALDIVision, a comprehensive bioinformatics tool for MALDI imaging, facilitates data processing, visualization and analysis of spatial distribution of individual ions in a tissue section. The program accepts imaging mass spectrometric data in Analyze 7.5 and imzML format files. It generates images in 2-D and 3-D modes. To correlate MALDI images with histological information of the sampled tissue, MALDIVision enables users to co-register and overlay a MALDI image over an optical image.